
NextGeM has innovative bio and digital technologies.We will expand our disease focusing areas, especially the areas with high unmet needs.
We will provide diverse novel values from
prevention to treatment by utilizing the
biotechnologies around hematopoietic stem cells.

Hematopoietic stem cell transplantation is a therapy by transplanting normal hematopoietic stem cells to patients with blood cancers or abnormalities of the blood and immune system and reconstructing the hematopoietic function.
The number of blood cancer patients eligible for HSCT is increasing not only in Japan but also worldwide. The reason why the incidence of blood cancers is soaring is because of the aging population.
HSCT is also used to treat intractable blood disorders such as aplastic anemia and severe combined immunodeficiency.
As a newer method, HSCT combined with gene editing technology is also expected to be applied to gene therapy. For example, in gene therapy using the ex vivo method (gene editing outside of the body), removing abnormal hematopoietic stem cells from patients with congenital diseases of the blood and immune systems from the body, correcting and normalizing it with gene editing technology, and re-transplanting into the patient for treatment.
Various problems have been pointed out, including donor shortage, burden to donors in collection, HLA (Human Leukocyte Antigen) mismatch, risk of side effects and complications, age and health limitations.. One of the challenges of HSCT is graft-versus-host disease (GVHD). This is one of the most life-threatening side effects of HSCT.
T cells, a type of donor-derived lymphocyte, contained in the HSC float, recognize the host’s cells as non-self and start an immune response (bio-defense), attacking the host’s normal cells. As a result, various disorders such as skin rash, diarrhea, and liver damage occur throughout the body, and severe cases are difficult to treat and can be fatal. The incidence of severe GVHD is reported to be approximately13% in HLA-matched unrelated transplants.
The removal of T cells is effective in preventing GVHD. Attempts to transplant CD34-positive cells only, which are reported to contain HSCs, to remove specific cells (TCRαβ cells) are being studied.
Recent advance in Hematopoietic stem cell transplantation. T Teshima. Japanese Journal of Transfusion and Cell Therapy, 66(1):3-6, 2020
Cord blood is an unlimited source of HSCs for allogeneic HSCT, however the amount of HSCs may not be sufficient for adults and the transplantation cannot be conducted. To solve this problem, it is necessary to develop culture technology that allows only HSCs to be amplified outside of the body. However, HSCs often differentiate into various cell types during culture, and as of 2020, no clinically applicable culture technology has been reported. (Based on internal survey)
Hoxb5 marks long-term haematopoietic stem cells and reveals a homogenous perivascular niche. Chen JY, Miyanishi M, Wang SK, Yamazaki S, Sinha R, Kao KS, Seita J, Sahoo D, Nakauchi H, Weissman IL. Nature. 2016 Feb 11;530(7589):223-7. doi: 10.1038/nature16943.
https://www.nature.com/articles/nature16943
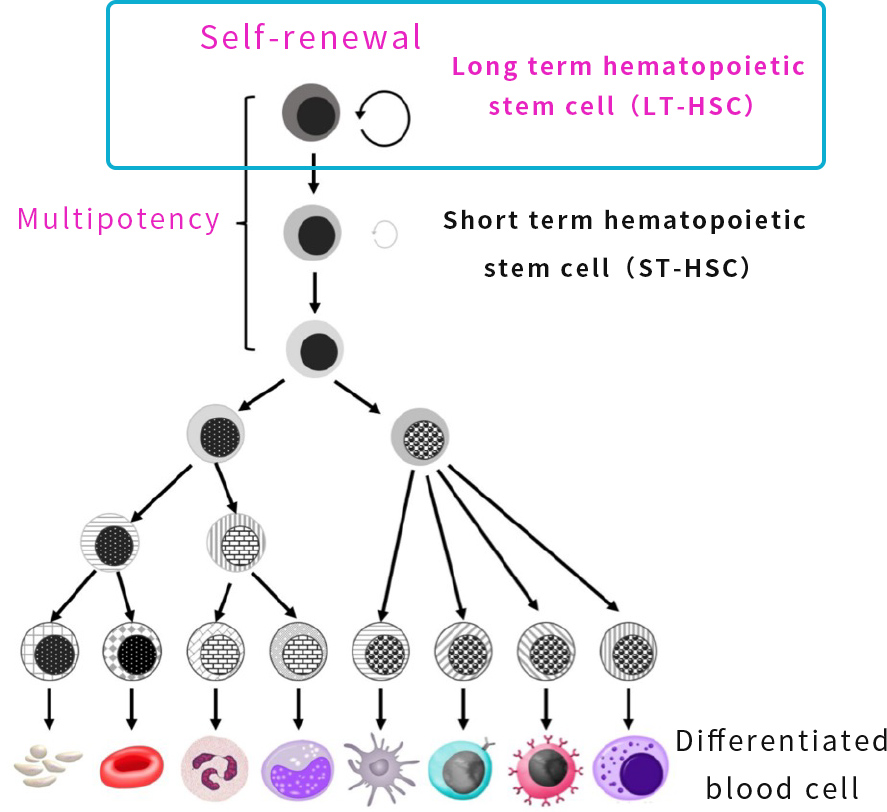
There are hundreds of biomarkers for human HSCs related to sustained hematopoiesis, and it is very difficult to selectively isolate human LT-HSCs from among them.
Therefore, by combining AI technology and in vitro functional experiments, we identify biomarkers that can purify human LT-HSCs from hundreds of biomarkers of human HSCs for sustained hematopoiesis and succeeded in concentrating LT-HSCs.
This technology can be applied not only to LT-HSCs but also to various other rare cells’ identification and isolation, and we are developing next-generation rare cell identification and isolation technology by integrating our proprietary technologies. This will be the fundamental technology which accelerates the understanding of many diseases, including precision medicine and rare and intractable diseases, and the development of therapies in the future.
Furthermore, we are developing transplantation methods for the following diseases by culturing and amplifying these concentrated LT-HSCs without differentiation even outside of the body using a culture medium developed by our company (click here for details).
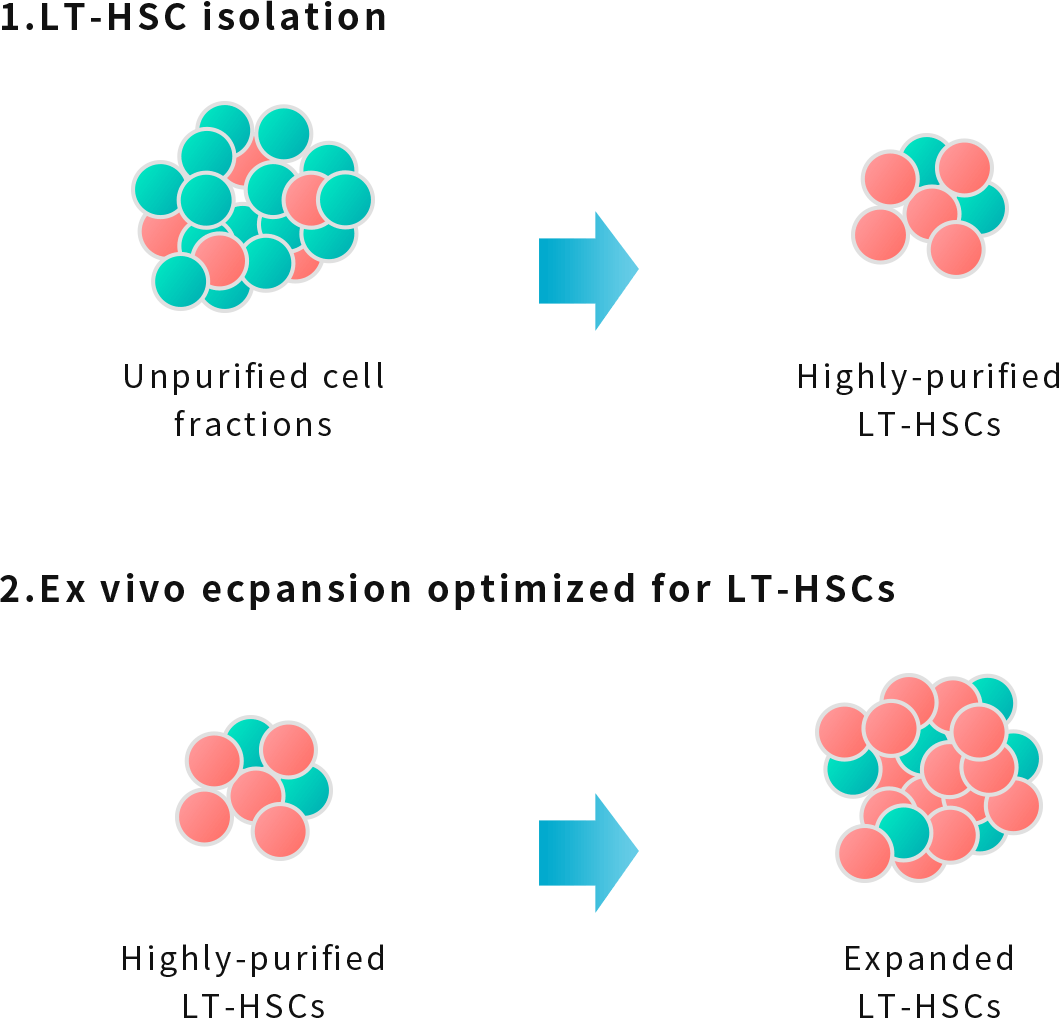
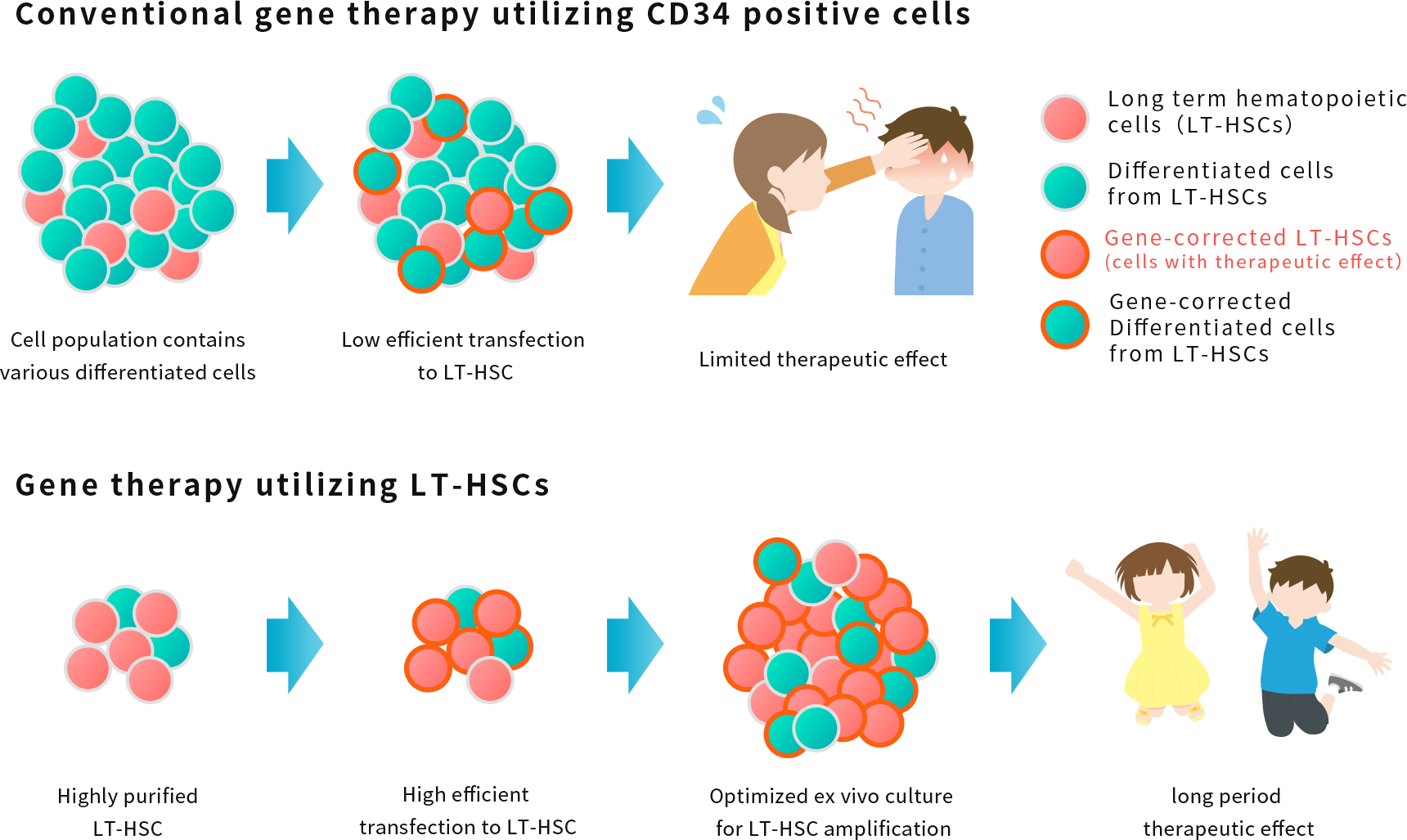
NextGeM is also developing gene therapy technology to extract congenitally abnormal hematopoietic stem cells from the body after isolating LT-HSCs and amplifying outside of the body using our proprietary technology, and then introduce the necessary genes into LT-HSCs and transplant back into the patient.
The target disease is expected to be intractable rare diseases in pediatric patients.
In addition, more than 6,000 types of genetic diseases (many of which are intractable rare diseases) have been reported so far. We aim to expand the scope of this technology to a wider range of intractable rare diseases through the development of this technology in the future.
As mentioned above, the number of HSCs that can be obtained from living bodies is extremely small, so in order for the widespread use of HSCs in the medical field, the cells need to be cultured and amplified outside of the body. However, under in vitro culture conditions, HSCs have the problem of differentiating into various types of blood cells as they proliferate.
Focusing on the fact that amino acids are important factors in the maintenance and amplification of undifferentiated HSCs*, NextGeM succeeded in developing a culture medium that amplifies HSCs including LT-HSCs while keeping them undifferentiated by adjusting the formulation. It is possible to maintain a higher percentage of myeloid progenitor cell-derived colonies (CFU-GEMM), which are the most undifferentiated cells at present, compared to conventional cell culture media.
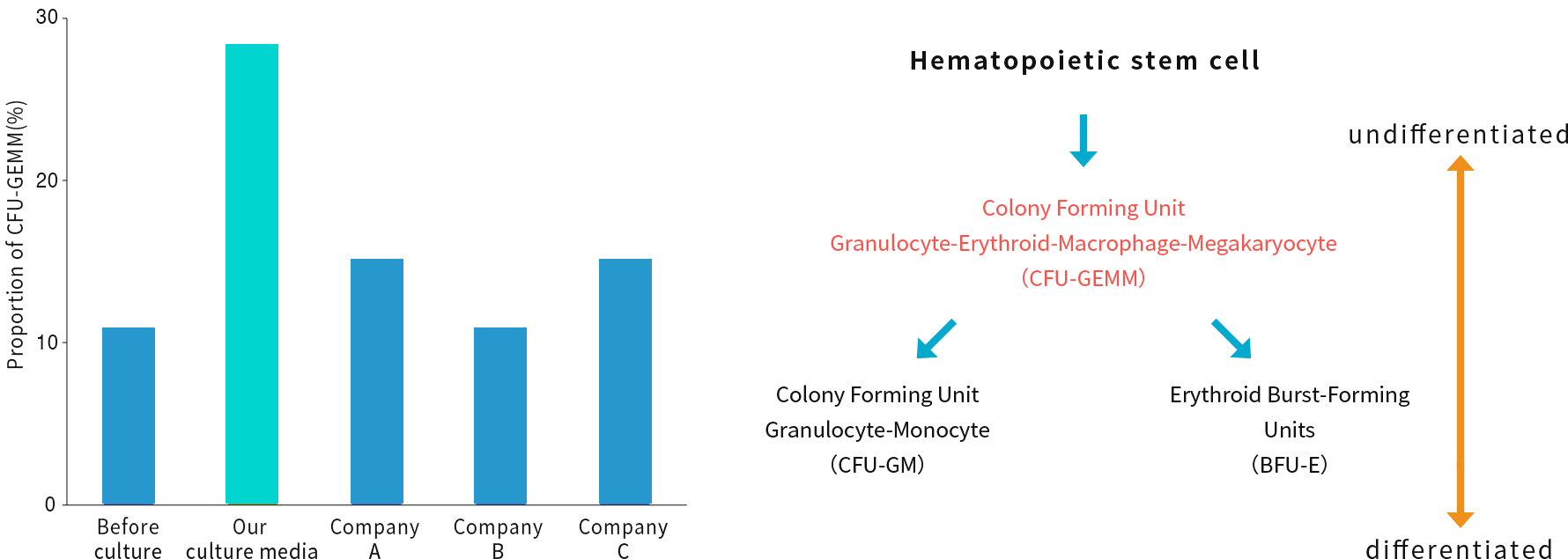
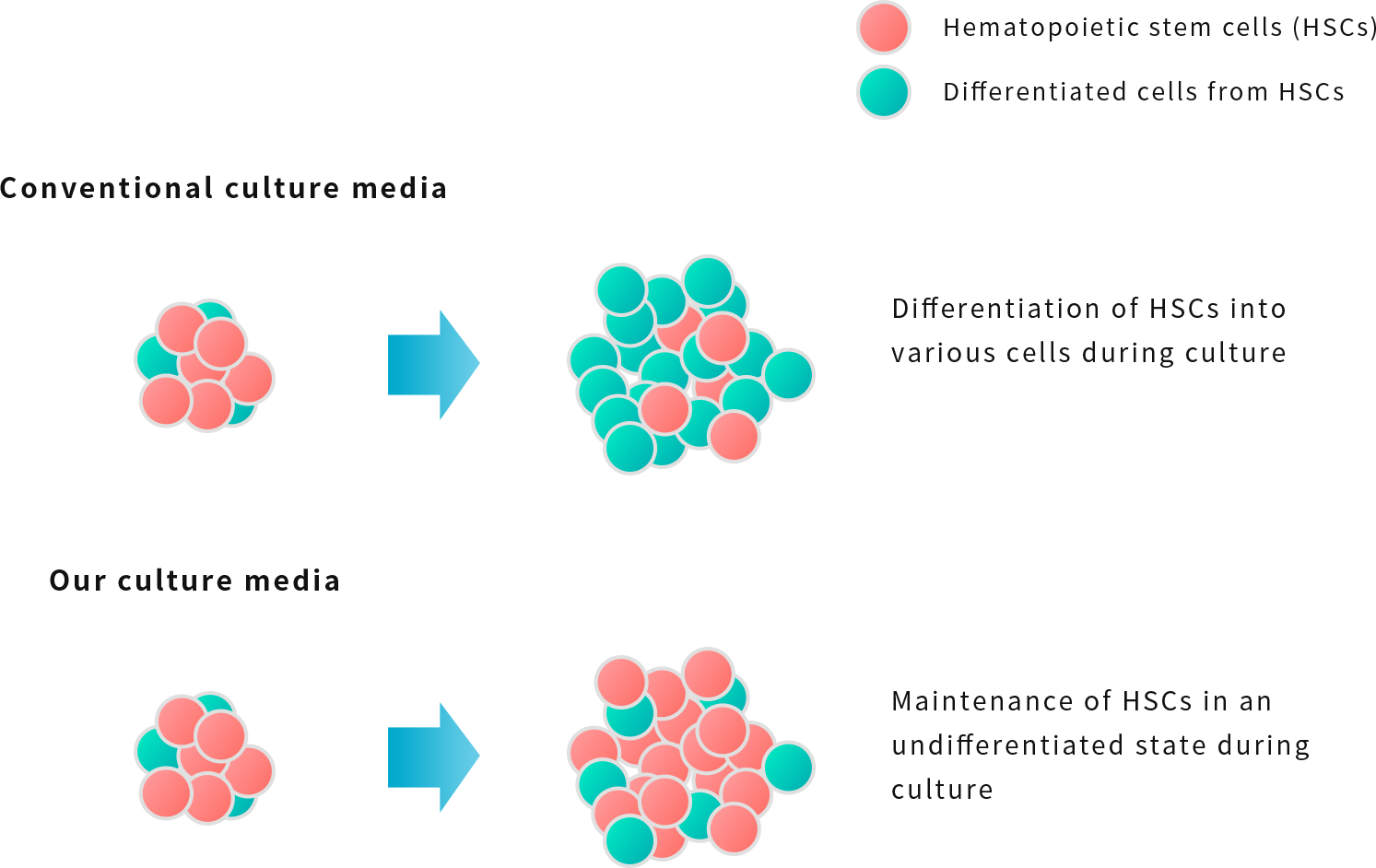
In addition, to reduce the risk of immune reaction after transplantation, we are currently developing a culture medium using plant-derived cell growth factors that do not contain animal serum. (Joint research with Mitsubishi Chemical Corporation)
NextGeM used CD34-positive blood cells and the hematopoietic colony forming unit assay (CFU assay) to evaluate HSC function after in vitro culture and optimized culture conditions that allow cell amplification while keeping undifferentiated HSCs. (Click here for details)
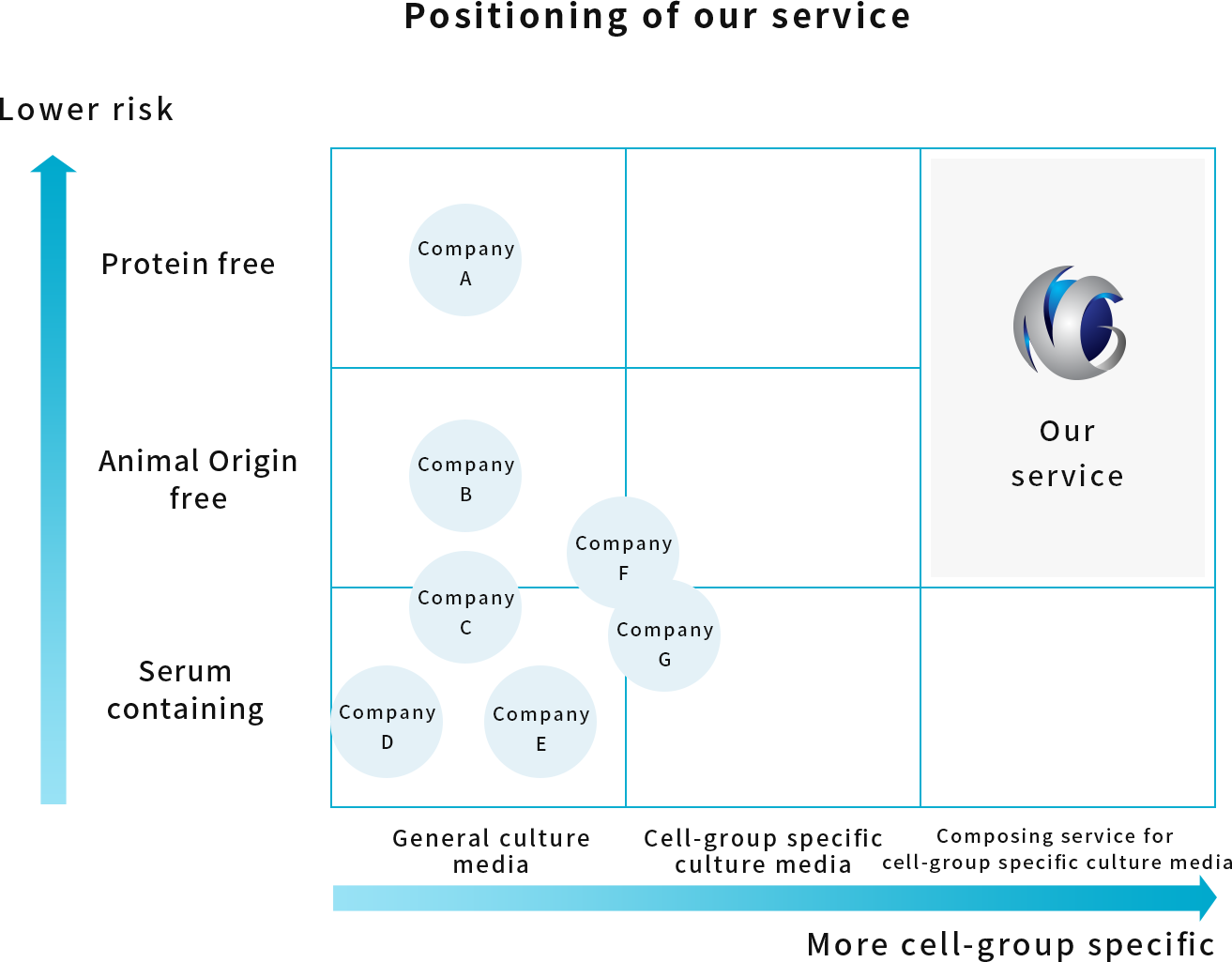
Utilizing these findings and experience in the formulation of components in HSC culture media and blood cell analysis techniques and know-how, we also offer services to develop the composition of culture media that will allow HSCs to differentiate into specific blood cell groups after isolation.
We aim to develop culture medium compositions that do not contain highly immunogenic serum, considering the medium’s transplantation use.
We are looking for partners to collaborate with our HSCs relevant technology in order to create diverse values in the following fields.
Expected collaborations (not limited to)
We are exploring various possibilities in addition to the above.
Let’s talk!
FiTTE has two functions around fertility treatment to solve clinical problems.
Despite a decrease in the fertility rate in Japan, the number of infants born with assisted reproductive technologies (ART) such as in vitro fertilization(IVF), Intracytoplasmic sperm injection(ICSI), cryopreservation of eggs and embryos, fresh embryo transfer, and frozen embryo transfer is increasing due to technological advances and later marriage. As of 2018, one out of every 16 newborns was ART infants. (Figure below)
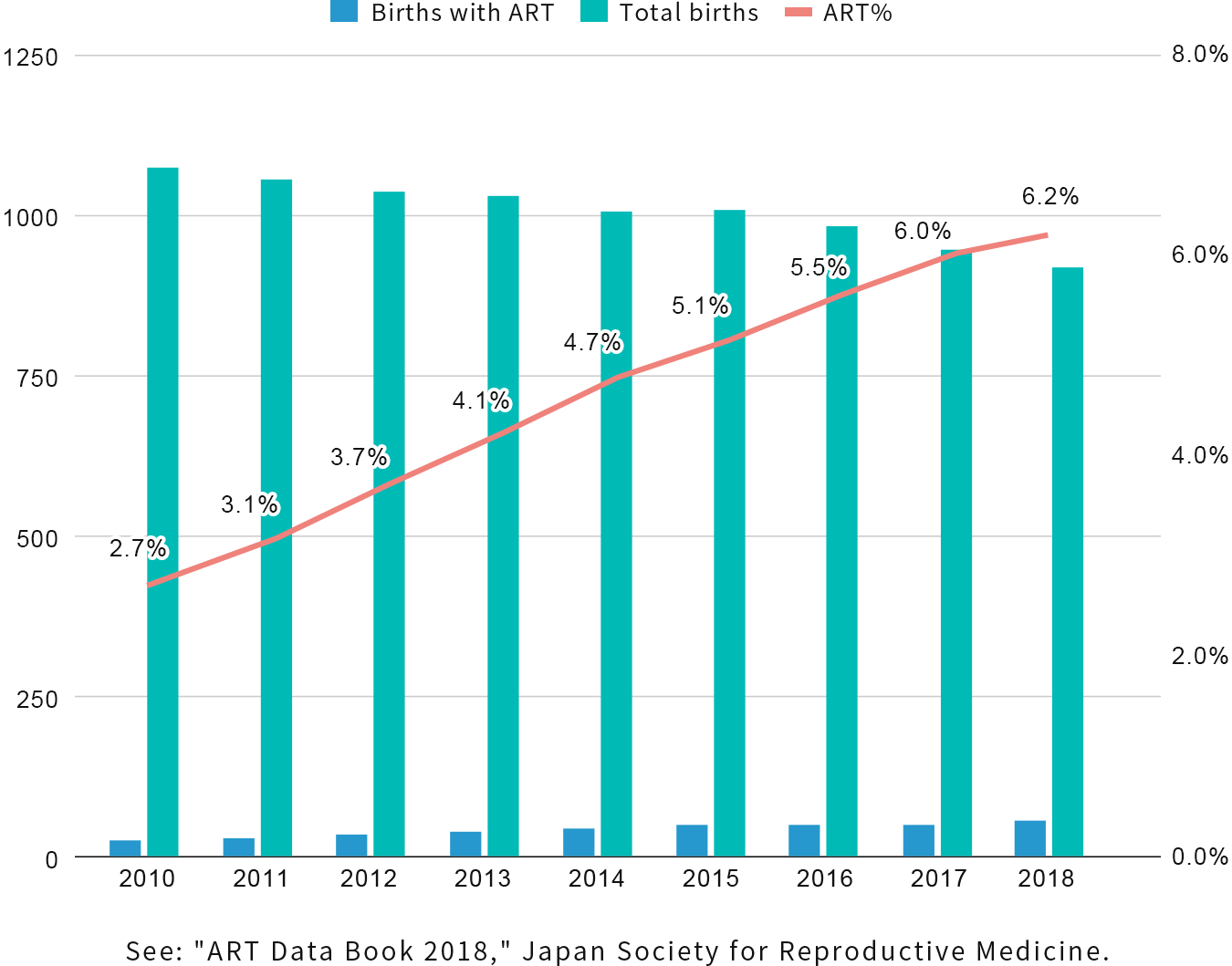
In Japan, where fertility treatment technology has advanced, infertility treatment has become so common and it is reported that one in every 5.5 couples experience it. However, infertility treatment is only partially covered by health insurance, and the financial burden on patients has been raised as an important issue.
Preimplantation genetic testing for aneuploidy (PGT-A) is a well-known testing method for embryos chromosomal aneuploidy, which is believed to be a cause of repeated miscarriages.
This method requires invasive procedures on the embryos, and there are concerns about its impact on pregnancy and fertility rates.
In addition, the cost of the procedure tends to be high, placing a heavy burden on the patient.

Based on the method (Ootsuki et al. Fertil Stertil. 2019) which determines the chromosomal aneuploidy non-invasively from the growth process characteristics of embryos (time point identification and area calculation) developed by the clinic and academic partners, NextGeM has developed the image segmentation AI to automatically screen the aneuploidy without human procedures. By conducting a retrospective observational study with approximately one hundred cases and tens of thousands of images in which
time-lapse videos were labeled by physicians and embryologists as training data, an image segmentation AI model was developed and enabled to extract embryo growth process features (time point identification and area calculation).
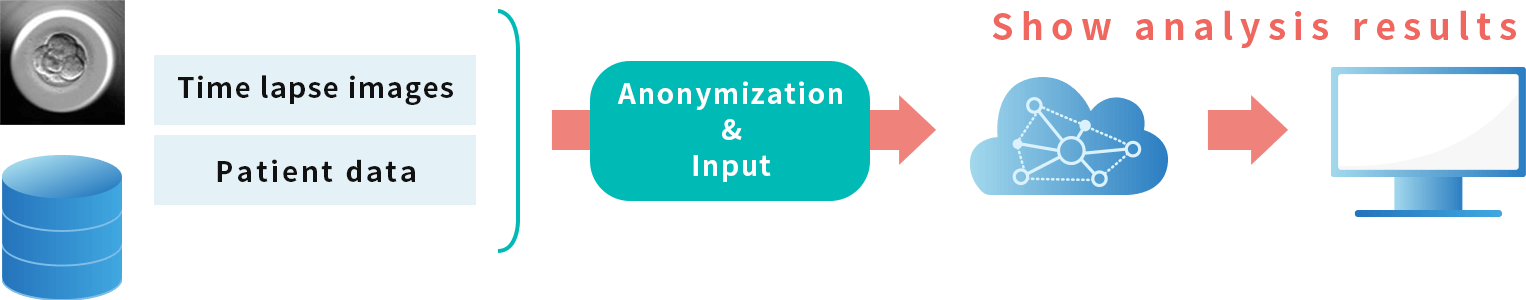
By uploading the time-lapse images of IVF/ICSI embryo culture into the software connected to the image segmentation AI placed on the cloud server, chromosomal aneuploidy determination information is provided to physicians.
This makes it possible to perform inexpensive, non-invasive, and rapid in-clinic testing and provide the necessary information.
(*Physicians should also check other clinical information and make a comprehensive decision.)
*NextGeM is conducting a prospective evaluation study with one of the largest (number of cases) infertility clinics in Japan.
[Supplemental Information]
A mechanism that enables anonymized processing within medical institutions has also been implemented.
A function that allows embryologists to correct the output results by the image segmentation AI is also under development.
Internet connection is required to download and the software can be used for research use.
For more details, please contact us by clicking on the link below
・Citations and references:https://www.fertstert.org/article/S0015-0282(19)30620-X/fulltext
In assisted reproductive technologies (ART), including IVF and ICSI, the condition of the embryo is considerably important for subsequent pregnancy and childbirth, but the current primary method is to visually evaluate the morphology of the embryo at the time of embryo transfer.
Various evaluation methods, such as embryo’s growth process analysis are being discussed at related academic societies, but there is still no established method to predict pregnancy and childbirth.
We have developed an image analysis AI that inputs time-lapse images of the embryo culture process and outputs the results of a five-stage evaluation of pregnancy and childbirth possibility.
As a retrospective observational study of approximately 20,000 cases, the image analysis AI was developed by extracting embryo growth process and morphological characteristics of embryos based on the correlation between time-lapse images of the embryo culture process and pregnancy and childbirth outcomes from the time-lapse images of embryos’ growth process.
(RMB 2022 ”A novel system based on artificial intelligence for predicting blastocyst viability and visualizing the explanation”)
Time-lapse images of IVF/ICSI embryo culture are uploaded into software connected to an image analysis AI placed in the cloud, and the results of a five-stage evaluation can be provided to physicians and
assist in the embryos’ selection for transfer.
・The software will assist physicians in embryo transfer (physicians should also check other clinical information and make a comprehensive decision).
・The image analysis AI model is placed in the cloud and linked to the software.
[Supplemental Information]
This analysis software is for research purposes.
A mechanism that enables anonymized processing data within a medical institution has already been implemented.
The software can be downloaded and used as software within the medical institution. Internet connection is required.
・Citations and references:https://onlinelibrary.wiley.com/doi/10.1002/rmb2.12443
This system facilitates matchings of donors and recipients in hematopoietic stem cells transplantation for malignant blood diseases.
When transplanting hematopoietic stem cells (HSC) for malignant blood diseases, serious side effects such as graft-versus-host disease (GVHD) may occur depending on the combination of donor and recipient.
However, it is difficult to select the best combination and predict prognosis because there is a vast amount of information on bone marrow donors, cord blood donors and peripheral blood donors and need years of clinical experience.It is difficult to select the best transplant source in many cases depending on pychisians’ clinical experience.
NextGeM developed a companion diagnostic system that enables the selection of the best donor candidate by simulating the prognosis for each individual donor candidate based on transplant donors’ information from cord blood banks and bone marrow banks and data and pretreatment information from patient or recipient.
We built a machine learning model that learns from approximately 2,500 patients at Kyoto University group hospitals over different time series. Compared with the existing COX proportional hazards model, it has achieved improved accuracy in predicting graft relapse-free survival(GRFS:GVHD-free, Relapse-Free Survival) after one year.
[Supplemental Information]
The system is deployed in a cloud server and is used as software within a medical institution. An internet connection is required.
・Citations and References
Published in Blood Advances “Establishment of a Predictive Model for GvHD-free, Relapse-free Survival after Allogeneic HSCT using Ensemble Learning「Establishment of a Predictive Model for GvHD-free, Relapse-free Survival after Allogeneic HSCT using Ensemble Learning」
This system supports diagnosis for myelodysplastic syndrome(MDS).
MDS (myelodysplastic syndrome) is a disease in which hematopoietic stem cells in the bone marrow become abnormal and stop producing normal blood cells.
The decrease in normal blood cells causes symptoms such as anemia, bleeding tendency, and fever associated with infection. Observed characteristics of the disease include “ineffective erythropoiesis ,” in which blood cells are destroyed in the process of becoming mature cells, and “dysplasia,” in which the formed blood cells are abnormally shaped. In addition, some patients may develop MDS or even “acute leukemia”.
In the diagnosis of MDS, a statistical report such as blood cell counts is used after clinical technicians observe bone marrow smear images, classify blood cells according to their morphological characteristics..
However, technicians need to observe approximately 500 smear images per patient and classify more than 30 cell types, which is very burdensome, and the classification accuracy depends on their experience.
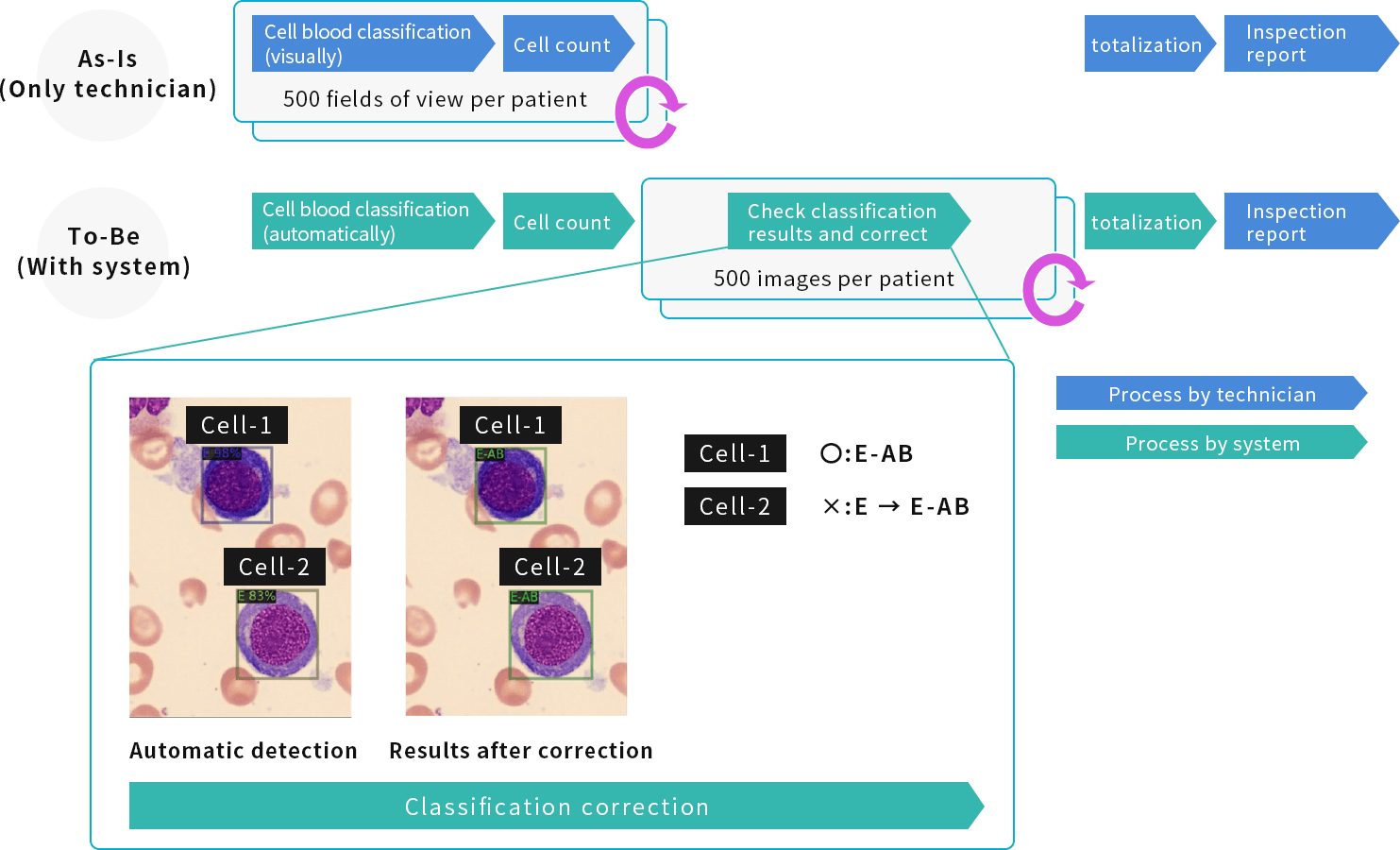
In order to speed up the classification and counting of blood cells by technicians, NextGeM developed an
image segmentation AI that automatically detects, classifies, and counts blood cells based on the input images taken from bone marrow smears. (*This program is not intended for diagnosis, treatment, or prevention of diseases.)
We trained the image segmentation AI to learn the morphological characteristics of blood cells, using blood cells labeled by physicians and technicians at Kyoto University Hospital as training data (approximately 100 cases and tens of thousands of images), .
Since this SaaS system is linked to the image segmentation AI model deployed in the cloud, it enables remote consultation with physicians and efficient information sharing between physicians and technicians who are separated even in the same hospital.
A function that allows technicians to correct the output results by the image segmentation AI is also under development.
This system supports diagnosis in bacterial infectious diseases.
Although it is important to accurately identify bacterial species and prescribe appropriate antimicrobial agents at an early stage of a bacterial infection, this is often not conducted because it is time-consuming to identify bacterial species by culture tests and so on without equipment (usually very
expensive) such as mass spectrometers. Hence, it relies on the experience of physicians and technicians (empiric therapy) and the antibiotics may be prescribed inappropriately. .
This lack of proper diagnosis and inappropriate prescription of antibiotics is one of the causes of antibiotics resistance (AMR: Antimicrobial Resistance).
BiTTE is an application that solves these problems by using AI image analysis technology to accurately estimate bacterial species, and by supporting the selection of appropriate antimicrobial agents in conjunction with antibiograms (antimicrobial susceptibility charts).
A smartphone is attached to an optical microscope via an attachment. Gram-stained images are captured with the smartphone’s camera. By uploading the image to the cloud, the results of the bacterial species estimation and the candidate antimicrobial agents to be prescribed are presented.
For the estimation of bacterial species, an image recognition AI model is built based on the gram-stained images of approximately 10,000 urine specimens from 1,000 cases and the bacterial species identification result of culture tests.
With regard to antibiotics suggestions, the system also displays information on spectrum scores, WHO’s AWaRe classification, and over-the-counter drugs information to enable
selection of more narrow-spectrum antibiotics.
Specimens are currently limited to urine, but will be expanded to blood and sputum in the future.
The system is also used in conjunction with medical equipment (under development) that automates the staining process in order to maintain consistent specimen staining quality and stabilize diagnostic imaging accuracy.
[supplement]
・Citations and References
Diagnosing bacterial infections using smartphones” Nature
本システムは、細菌感染症における診断上の課題を解決し、世界的にも注目されている薬剤耐性問題の解決を目指します。
As exemplified by the ‘Action Plan for Antimicrobial Resistance (AMR)’ formulated by the Ministry of Health, Labor and Welfare (MHLW) in Japan in 2016, countries around the world are also taking steps to combat AMR.
However, in emerging countries, due to the shortage and uneven regional distribution of medical specialists, and inadequate medical platforms, diagnosis and treatment, including identification of causative organisms by gram staining and selection of appropriate antimicrobial agents, have not been properly carried out.
We have created a remote diagnostic support platform as a solution for regions such as these emerging countries, where there is a lack of infectious disease specialists or a lack of medical platforms.
This platform is a service that allows primary care physicians, who are having troubles in diagnosing infectious diseases, to upload medical images to the cloud and request an identification support by a medical specialist anywhere in the world.
Multiple images can be uploaded for each patient and specimen.
Fees are based on a pay-per-read request basis, and the reading response time can be determined.
The platform also has functions to request a quick response and to request cross-checking by multiple experts.
We are looking for partners who aim to create diverse value by collaborating with the following technologies related to AI and SaaS that we possess.

Collaborations envisioned (not limited to)
We are exploring various possibilities in addition to the above.
Let’s talk!